Back to Tutorial pages
- Scalar: a single value
- Vector: a column of values - one value of each node (vertex) in the mesh.
GFtbox Growth factor naming conventions
Prefix
| id_ |
identity factors (non-diffusible). Identity factors can be labelled as on the A or B side (ida_, idb_)
|
| s_ |
signalling factors (diffuse). Polariser level is unique and has a particular name P.
|
| v_ |
visualising labels (e.g. theta, bend, kpar, other labels)
|
| f_ |
finite element manipulation (e.g. seams, making holes)
|
Suffix
| _p |
vector representing promotor: how much the expression of the factor is being promoted in the model at every node (vertex) in the mesh.
|
| _l |
vector representing the level of morphogen, i.e. *_p times *_a at every node in the mesh. This gets updated at the end of the interaction function and might change according to growth or mutation, see *_a.
|
| _i |
scalar index into the main data structure morphogen field, m.morphogen.
|
| _a |
scalar specifying activity: by default it is 1 and if the factor is mutated or silenced then it is 0. However, it can be in the range [0,...,1].
|
Standard morphogens
Specifying growth
| kapar_p, kapar_a, kapar_l |
vector that specifies growth on the 'A' side parallel to the polariser and in the plane of the tissue at every node.
|
| kbpar_p, kbpar_a, kbpar_l |
vector that specifies growth on the 'B' side parallel to the polariser and in the plane of the tissue at every node.
|
| kaper_p, kaper_a, kaper_l |
vector that specifies growth on the 'A' side perpendicular to the polariser and in the plane of the tissue at every node.
|
| kbper_p, kbper_a, kbper_l |
vector that specifies growth on the 'B' side perpendicular to the polariser and in the plane of the tissue at every node.
|
| knor_p, knor_a, knor_l |
vector that specifies growth normal to the plane of the tissue (i.e. thickness) at every node.
|
Controlling growth
| P |
vector that specifies Polariser level at every node.
|
| strainret_p, strainret_a, strainret_l |
vector that specifies strain retention at every node. It is a number between 0 (no strain retention, default) and 1 (full strain retention).
|
| arrest_p,arrest_a,arrest_l |
vector that specifies an overall inhibition of growth at every node of the mesh. It is a number between 0 (no overall inhibition of growth, default) and 1 (fully inhibited).
|
User (your) morphogens
Within the interaction function all the usual Matlab variables and types are available. Variables that are to be visible from the GUI must follow certain conventions and be declared within the GUI, see below.
(Note to programmers: we could have produced a class library to handle the different types of variable - identity factors, signals, etc. However, in our view it is too early in the development of software for modelling the growth of biological tissues to be confident of what will be required. Too build a class library now would be to build a conceptual straightjacket that would inhibit further research.)
Identity factors, signalling factors, viewable variables
Growth is all about patterns of growth factors interacting in space (spatially) and over time (temporally). It is, therefore, essential to be able to view them on the mesh. This is the main purpose of the graphical user interface (GUI). Such variables are created (declared) from the GUI as follows. They have a name e.g. id_plusorg. This is how the variable will be known as a case insensitive string (according to Matlab conventions, 'id_plusorg' or 'ID_PLUSORG'). Within the interaction function this name will be lowercase and have a suffix e.g. id_plusorg_p - see above.
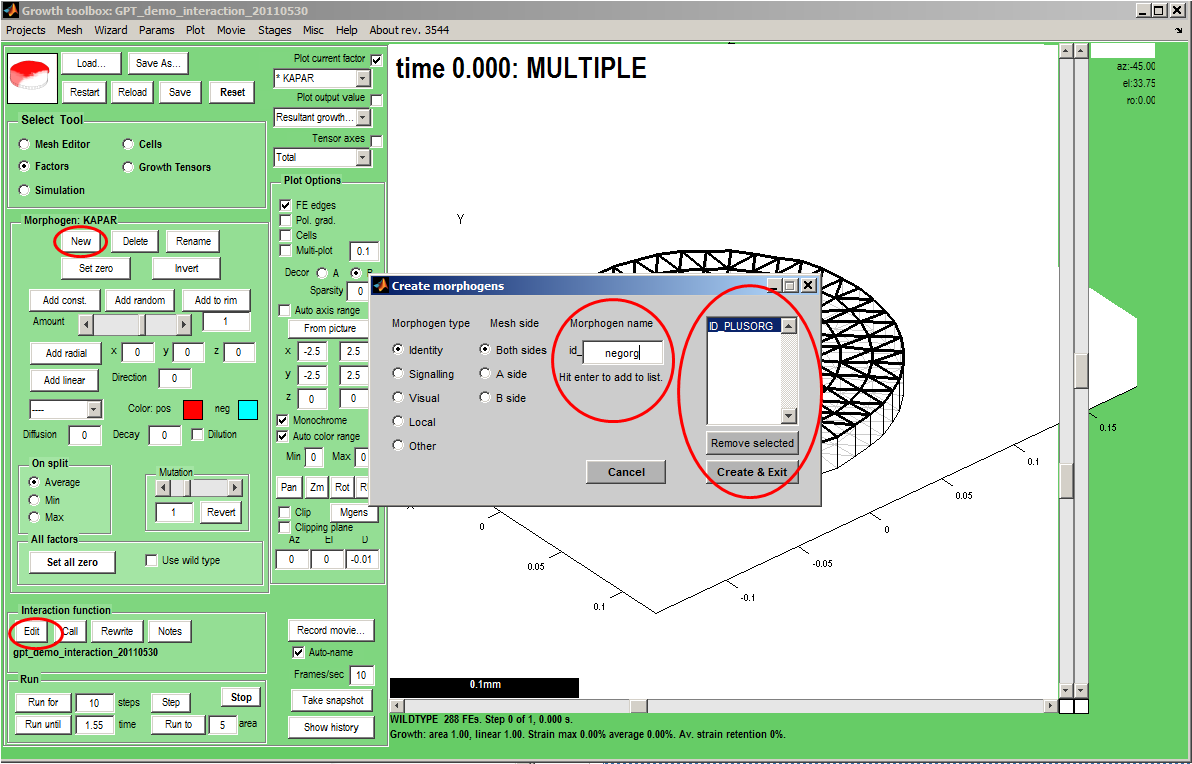
Adding (declaring) variables that are visible in the GUI. Select Tool:Factors makes the Morphogen Panel visible. We add a two new morphogens, id_plusorg and id_negorg. Select a type (in this case id_), key in the name, press Enter and the name will be added to a list. Finally, create the morphogens and exit the dialogue. Select a colour for each, Panel: Morphogen:Color. pos. and Color. neg (for negative values).
Save to make sure they are saved in the Mesh. Pressing Panel:Interaction Function:Edit will reload the editor with a revised copy of the interaction function. The new copy will retain any edits that you have made already together with the new variables.
|
|
